- Consortium Research Platform on Genomics (CRP-Genomics)
Clustergene DB for easy retrieval of unigenes and microsatellite markers in Cluster bean (www.nrcpb.res.in)
Cluster bean is an important vegetable, forage, and industrial crop. It is a drought tolerant annual legume that can grow well in marginal lands but still respond to input agriculture. India is the largest producer of cluster bean accounting for 80% of its production.
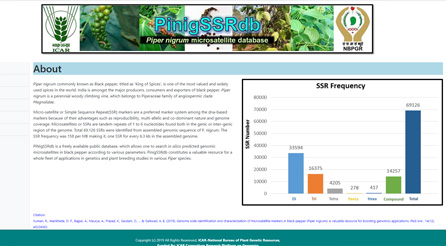
PiNigSSRdb: Database of microsatellite markers in Piper nigrum (black pepper):
PiNigSSRdb is a freely available public database, which allows one to search in silico predicted genomic microsatellites in black pepper according to various parameters. PinigSSRdb constitutes a valuable resource for a whole fleet of applications in genetics and plant breeding studies in various Piper species. Total 69,126 SSRs were identified from assembled genomic sequence of P. nigrum. The SSR frequency was 158 per MB making it, one SSR for every 6.3 kb in the assembled genome.
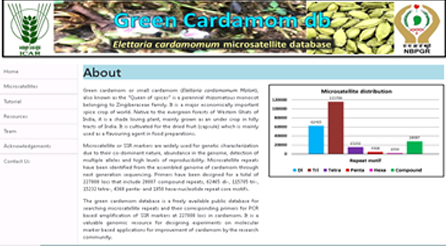
ElCarSSR db: Database of microsatellite markers in Eletaria cardamom (green cardamom)
Elettaria cardamomum microsatellite database (CardamomSSRdb) is an interactive and relational online database that contains comprehensive information on small cardamom genomic SSRs that were identified using MISA script and primers designed using Primer 3 software. Other than genomic SSRs data and statistics, the database also contains step by step pictorial tutorial to facilitate hassle free usage by the user
The database can be accessed at http://www.nbpgr.ernet.in:9092/

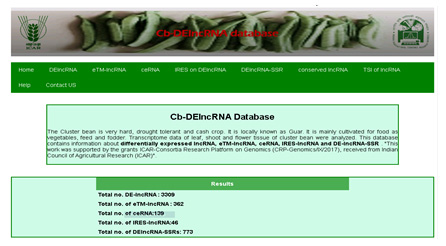
CbLncRNAdb (Cluster Bean LncRNA Database)
CbLncRNAdb provides information on lncRNAs, SSRs, miRNAs, lncRNAs bearing SSRs, miRNA bearing SSRs and endogenous target mimic (eTMs) lncRNAs. Search facility for lncRNAs and Highly Probable lncRNAs (HPlncRNAs) is given in the database. Statistics menu is also provided to get summary information of the database.
Database of Hilsa Transcriptome and SSRs
Hilsa Transcript SSR db, a searchable database, based on transcritptomes of Hilsa shad from five tissues, muscle, kidney, liver, testis and ovary and simple sequence repeats present in the transcripts.
The database can be accessed at https://www.nbfgr.res.in:802/index.aspx

The Cluster bean is very hard, drought tolerant and cash crop. It is locally known as Guar. It is mainly cultivated for food as vegetables, feed and fodder. Transcriptome data of leaf, shoot and flower tissue of cluster bean were analysed. This database contains information about differentially expressed lncRNA, eTM-lncRNA, ceRNA, IRES-lncRNA and DE-lncRNA-SSR. "
The database can be accessed at hhttp://backlin.cabgrid.res.in/Cb-DElncRNA/
SNP Search Tool: The dbVAST (database for variations associated with shrimp transcripts), an on-line tool to browse single nucleotide polymorphisms in annotated transcripts of two important shrimp species, Penaeus vannamei and Penaeus indicus has been developed.

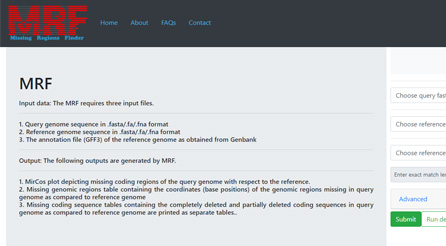
Find missing regions in viral genomes with MRF: A web-tool namely MRF (missing region finder) has been developed to find missing coding sequences in viral genomes when compared to a reference genome. The MRF has advantages in conducting comparative genomics studies in pathogens with unique genome sequences.