ICAR-Indian Agricultural Research Institute (IARI), New Delhi
Microbial plant pathogens are major threat to agricultural production worldwide. Agriculturally adapted and rapidly evolving pathogens belong to all major taxa especially fungi, virus and bacteria has the potential to reduce the genetic capability of crop plant. In order to sustain agricultural production, a sound and effective disease mitigation strategy must be in place. With the increasing global connectivity, threat of invasive pathogen and exotic diseases affecting our crop production is quite a reality. Rapid advancement in genomics technology in the recent years has altered the landscape of plant pathology research once for all. Therefore, it is hoped that the modern genomics technologies have the potential to assist the development of new strategies for crop protection.
Objective
- To develop pathogenome browsers for nationally important plant pathogens, including Tilletia species, Bipolaris species, Ralstonia, and Magnaporthe species, which significantly impact agricultural and horticultural crops.
- To create high-throughput array-based genomic chips for the rapid monitoring and detection of nationally relevant plant pathogens.
- To identify novel inhibitors or detoxifying agents for pathogen sporulation based on transcriptomics data generated from these pathogens.
- To identify host-specific transcripts involved in pathogenesis using transcriptomics data.
- To enhance expertise and human resources in pathogen genomics through workshops and training programs.
Research targets/activities
Structural and functional genomics interatomic with host
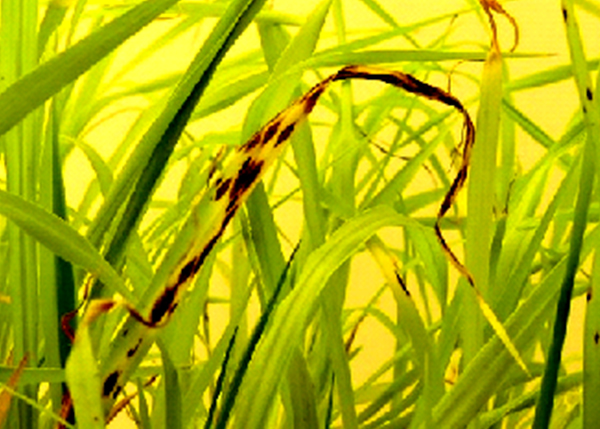
- Magnaporthe infecting cereals (rice, pearl millet, finger millet)
- Tilletia infecting wheat.
- Ralstonia infecting vegetables.
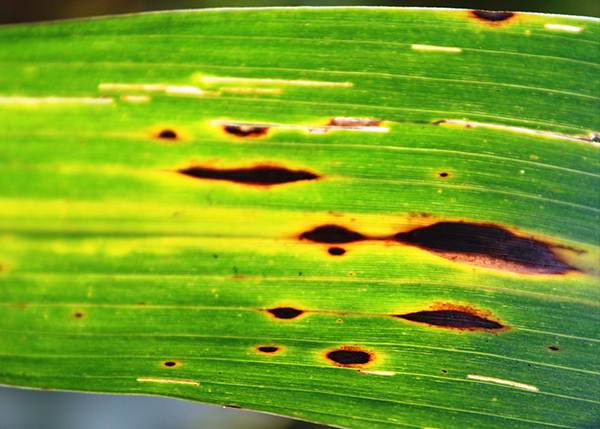
- Bipolaris infecting cereals
- Development of a diagnostic assay for the detection of Bipolaris sorokiniana based on the genome sequence